SARS-CoV-2 Multisite analysis
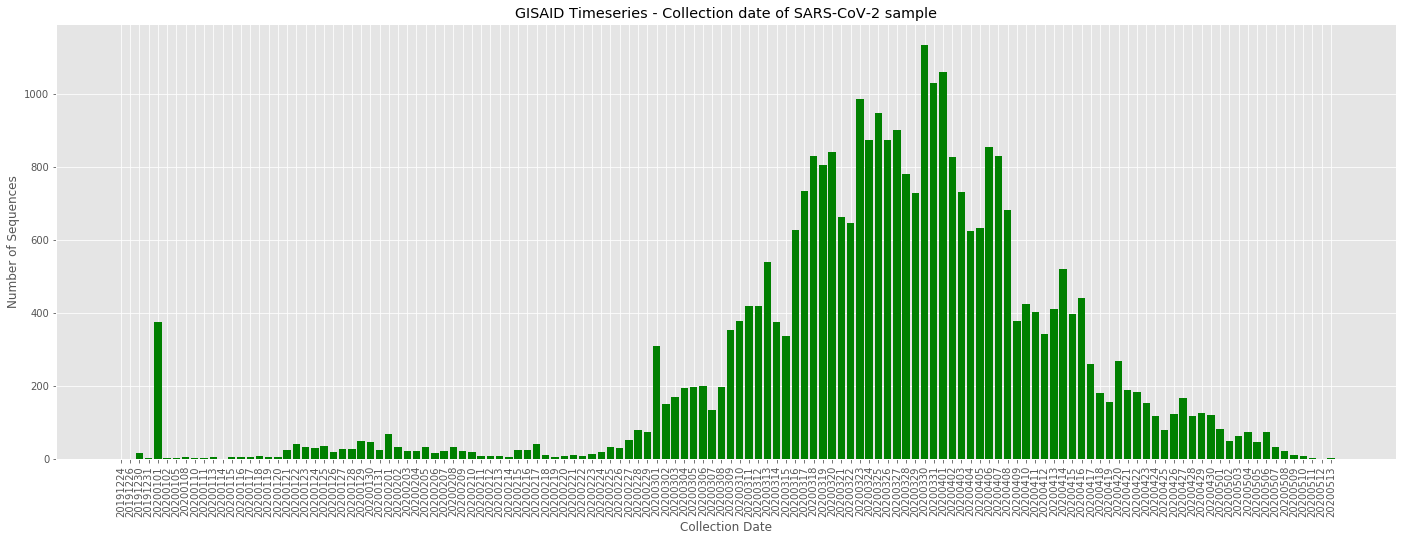
This analysis was constructed to look across "interesting" sites which show evidence to be acting under natural selection. {Link to observable}. Daily pulldowns from the GISAID portal were used. These SARS-CoV-2 sequences were codon-aware aligned and the following genes have been critiqued (S, M, N, ORF1a, ORF3a, ORF7a, ORF8). From the "compressed" alignments, the site number and codon at that position were recorded for all available sequences from that day. Time-series analysis was performed in a manner where each day represents the cumulative dataset of information pulled down from GISAID. Using this site/codon "Code", a haplotype or mutation sequence / sequence fingerprint was created. The graphs below describe the observed frequency of that genes "haplotype" on that day.
| Gene | SARS-CoV-2 "Interesting" Sites |
| M | 30,69,109,132,141,143 |
| N | 13,20,24,55,67,108,119,139,144,151,173,207,208,210,218,253,291,305,318,360,377,393,397,417 |
| ORF1a | 16,84,86,92,166,210,265,274,286,333,398,428,454,447,486,519,549,580,609,650,652,717,748,872,892,894,924,1100, 1143,1158,1160,1168,1202,1208,1250,1330,1352,1415,1640,1822,1840,1921,2062,2067,2098,2124,2144,2153,2242,2249, 2391,2405,2470,2497,2500,2557,2575,2635,2796,2839,2842,3027,3058,3070,3071,3072,3076,3090,3227,3278,3308,3323, 3353,3357,3447,3483,3512,3542,3580,3603,3606,3639,3644,3714,3725,3755,3757,3777,3856,3858,3870,3884,3899,3958, 3983,4023,4097,4177,4220,4379,4381,4389 |
| ORF3a | 42,43,57,68,99,125,131,240 |
| ORF7a | 70,92,121 |
| ORF8 | 8,84,121 |
| S | 29,61,138,231,294,598,614,620,623,675,723,731,824,934,936,1044,1100 |